
Have you ever been tempted by the ads for 23 and Me, a DNA testing service so named as each human has 23 chromosome pairs? Or maybe you have already taken the plunge and received your results. Millions have used this test or another to learn their ethnic identities, investigate their health risks, and connect with cousins close and distant (Ossola, 2015). Or maybe you have used a DNA test to identify the breeds that make up your adopted rescue dog, or to verify you have purchased a purebred canine pal. In recent years the uses and potential uses of DNA profiles have exploded. DNA tests are now widely available for deadly serious reasons—like identifying the likely physical traits of the perpetrator of a crime based on a DNA sample found at the crime scene—and for the just-plain curious. While the media has made human DNA testing well known, less may be known or understood about livestock applications.
Suppose you are a cattle rancher, feedlot operator, restaurateur, lover of a fine steak, or just someone wanting to include more beef in family meals—is there a DNA test for beef cattle that can benefit you? Progress has been bumpy, but DNA tests for beef cattle are on the market and beginning to take hold with mainstream cattle producers. Current and prospective tests rely on underlying investments in genomics science and its applications to beef cattle, a research priority for the U.S. Department of Agriculture (USDA) for a number of recent years. Between 2008 and 2014 the USDA awarded $61.5 million for studies of beef or cattle genomics [based on a STAR METRICS search on “(genomics OR genetics) AND (beef OR cattle) NOT dairy (OSTP, 2016)." It is not too soon to contemplate the economic implications and returns to these investments. What information is provided by current DNA tests for cattle; and what might be provided in the future? How widespread will adoption be, and how will adopters use test results? Will those decisions lead to a different quality product; to more affordable beef; to a restructuring of the beef industry? How will the benefits be shared in the multi-stage beef value chain?
Most DNA tests—for yourself, your dog, or your cow—now use single-nucleotide polymorphisms (SNPs), pronounced “snips”, to identify genetic variations or similarities among individuals, or between an individual and a reference group. What are these SNPs?
The DNA molecule consists of a string of four chemical “bases,” adenosine (A), thymidine (T), guanine (G), and cytosine (C). To form the characteristic DNA double helix, hydrogen bonds form between the paired bases, A with T and G with C. A gene is a specific string of these paired bases that provides instructions for making or regulating a particular product, such as a hormone or enzyme. The order of the bases on a string is called a sequence, so the purpose of sequencing a genome—or partial stretch of a genome—is to reveal the linear order of the four bases. Sequencing can then be used to identify places where there is a substitution of one base for another in some percentage of the population being studied. For example, along a particular stretch of the genome researchers might find a G in a particular position 72% of the time and an A 28% of the time. These places, where a commonly located base is replaced with a different base, are the SNPs—also sometimes referred to as point mutations. These SNPs—which may be positioned in a gene or in the DNA sequences in between genes—can serve as markers for diseases and other inherited or genetically influenced traits that can be passed along to offspring. Genomics researchers look for correlations between SNPs and traits of interest, and evaluate the statistical validity and strength of the observed relationships.
Daisy, or any other cow, for that matter, has approximately 22,000 genes arranged on 60 chromosomes—or 30 pairs, one of each pair from each parent. The large number of possible orderings of the four bases that compose these many thousands of genes and DNA sequences in between genes means there are multiple millions of possible SNPs. In fact, the b. taurus cattle genome contains about 4 million SNPs (Seidel, 2009). In 2009—the same year Science magazine reported the first sequencing of the bovine genome (The Bovine HapMap Consortium, 2009)—the life sciences company Illumina released a commercial SNP evaluation system for cattle that made it possible to rapidly analyze up to 50,000 SNPs spaced throughout the genome. This “50K chip,” which was developed in collaboration with USDA’s Agricultural Research Service (ARS), Pfizer Animal Genetics, University of Missouri, the National Association of Livestock and Artificial Insemination Cooperatives in France (UNCEIA), and the French National Institute for Agricultural Research (INRA), provided a substantial boost to research aimed at matching SNP profiles with economically important cattle traits (Matukumalli et al., 2009). As an indicator of the boost received, the number of USDA-supported project progress reports that include the terms “SNPs” and “beef cattle” rose from 14 in 2000 to 90 in 2012 (USDA-NIFA, 2016b). Recently, the process of identifying associations between individual SNPs and traits of interest has been replaced by a process known as genomic selection, which looks at the relationships between all 50K or more SNPs imputed up through statistical inference to the full sequence (Meuwissen, Hayes, and Goddard, 2016).
Animal scientists conducting SNP association studies face a number of significant challenges despite huge advances in genomic prediction technology. First, there must be a sufficient number of “phenotypes” to match with SNP profiles. Phenotypes are actual cattle that exhibit objective measurements of the economically important traits of interest. Many hundreds or thousands of animals may be needed for statistically significant correlation results. Some breed associations, such as the American Angus Association, have long collected extensive amounts of information on offspring of sires, which makes SNP correlation studies more feasible for Angus cattle than for other less established or less-studied breeds. Public research entities, such as USDA’s Meat Animal Research Center (MARC), establish study populations for other breeds and mixed breeds; MARC validates study results by comparing them with results of SNP studies conducted at other research centers with other large cattle reference populations, for example in Australia and Canada (Pollak, 2012).
Second, even for breeds with large amounts of phenotypic data available for SNP matching studies, data may not have been collected for all genetic traits or trait indicators of interest. For example, a cattle breed association might have recorded information on birth, weaning, and yearling weight, but not on feed conversion efficiency, meat tenderness, nutritional quality, or a newly emerging disease. Feed conversion efficiency, for example, is an especially costly trait to study in cattle and therefore not routinely measured by breed associations.
Third, traits of interest in the cattle sector have varying degrees of “heritability,” which refers to the proportion of phenotypic variation due to genetic variation. Most traits are influenced by a combination of genetics and environment, just as in humans. For example, certain indicators of reproductive performance in cattle, such as scrotal circumference, have high heritability and others, such as calving intervals, have low heritability (Field, 2007). The degree of heritability affects the degree of confidence with which scientists can link SNPs with inherited performance traits. Furthermore, no matter how strong the match between an animal’s SNP profile and its propensity to pass along desired traits to its offspring, it still “takes two to tango.” The other parent’s genomic profile may be just as important in determining the performance merit of their calves.
Genomics research for beef cattle has tried to match SNPs with both simple and complex traits. Simple traits are those controlled by a single pair of genes. Simple cattle traits include qualitative—or observable—traits such as coat color, pigmentation, horned or not horned, double muscling, and a variety of undesirable genetic defects and diseases that can plague cattle producers, such as dwarfism, hairlessness, marble bone disease, mulefoot, and palate-pastern syndrome (Field, 2007).
Some simple traits have significant economic benefits or costs. Returns to genetic technology designed to select for hornless animals were found to be quite high in Australia where a dominant breed is the naturally horned Brahman (Griffith and Burrow, 2015). The simple trait “color” may be economically valuable for some cattle producers. The Certified Angus Beef program, for example, requires the animal be at least 51% black to meet the phenotype requirement, and Angus steers and heifers may bring higher prices than non-Angus contemporaries (AgriCultured, 2016). Double muscling—the result of a SNP in the myostatin gene that controls muscle development—can lead, according to some, to a leaner and healthier choice for consumers, but also to larger calves and more calving difficulties (Alford et al., 2009; Schaffer, 2015).
However, most economically relevant traits in cattle are complex traits, which means they are controlled by multiple pairs of genes, and are therefore more difficult to match with SNPs than are simple traits. Complex traits are typically quantitative—or measurable—traits that may be grouped into those that affect productivity, such as various dimensions of reproductive performance and weight gain; production costs, such as feed conversion efficiency; or product quality, such as meat tenderness and marbling. Genomic scientists search for and have found SNP associations for quite a few of these elusive economically relevant, complex traits, but with varying degrees of statistical validity. As just two examples among a great many, one study of feed conversion efficiency found strong SNP associations on seven different chromosomes (Abo-Ismail et al., 2014); and a study of calving ease found 13 closely associated SNPs on just one chromosome (Bongiorni et al., 2012). However, complex traits typically have low heritability, such that parental DNA alone is insufficient for predicting phenotypes because other factors, such as temporary or permanent environmental influences, genomic imprinting, and genetic recombination, come into play.
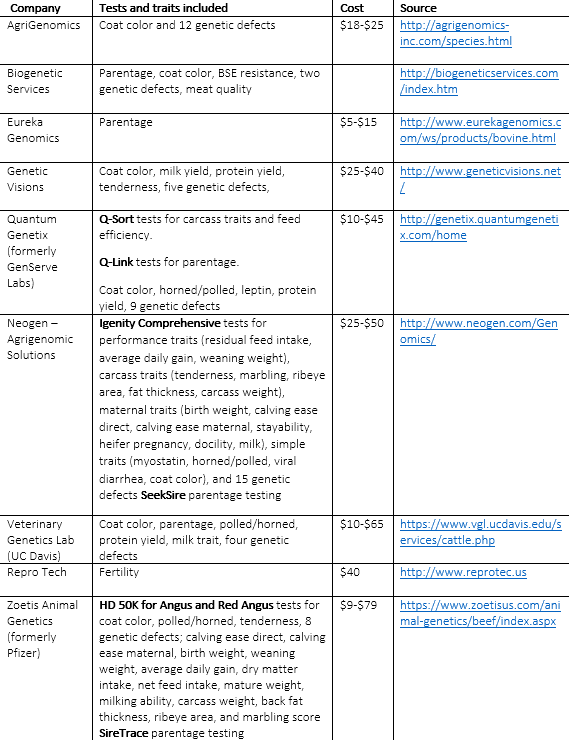
Source: Van Eenennaam (2010) and author updates.
There is no DNA test called 30 and Daisy, although we think there should be. However, as shown in Table 1, there are a number of genomic tools for beef cattle available on the market now. Some earlier tests were removed from the market due to weak predictive performance, and the genomic testing industry is still experiencing considerable flux—as evidenced by oft-changing company names and products. The currently available tests typically offer results for simple traits, as well as parentage verification. Only two companies—Neogen and Zoetis (formerly Pfizer)—offer comprehensive genomic prediction tools at prices ranging from less than $10 to test for parentage and some simple traits, to $79 to obtain information on a panel of economically relevant complex traits.
The cattle industry has been selecting and breeding for genetics for over 200 years (Olmstead and Rhode, 2008). Genomic tools, if effective, “merely” speed up the process by providing information about inherited traits that are not readily observed or cost-effectively measured, and about the expected genetic merit of breeding bulls and cows that are too young to have proven performance records. What each cattle producer or buyer has to decide is if the current cost of buying a genomic tool is worth the benefit down the road in the form of more valuable or less costly-to-raise offspring. This calculation depends on both how valid the test is—how accurately it can predict the desired traits in the offspring—and how much value there is in enhancing those traits in the offspring.
Who are these cattle producers and buyers who might want to consider benefits v. costs of DNA testing? Beef production involves multiple stages along a vertical value chain: a seed stock sector, cow-calf operations, stockers and feedlots, processors, retailers, and final consumers (Field, 2007). In relation to poultry, hogs, and dairy, beef production involves longer biological cycles spread across a larger geographic area. Additionally, ownership of the animals in the value chain can change multiple times across the industry segments (Mintert, Shcroeder, Brester, and Feuz, 1997). Consequently, there is less vertical coordination and control in the beef industry than in other livestock sectors. In a practical sense, this has implications for how benefits of using genomic tools are likely to be captured or shared in the beef supply chain. For example, a typical Wyoming cow-calf operator does not retain ownership of her animals once they’re loaded on a truck and bound for feedlots, usually in neighboring states. Could she recoup the costs of having invested in DNA tests? Norwood and Lusk (2008) and Van Eenennaam and Drake (2012) make the case that although investments in beef genomic tools may generate value for each sector of the beef industry, they will be less profitable than in poultry, hogs, and dairy because of the relative lack of vertical control in the beef sector.
What is actually known about current and prospective adoption and economic returns to DNA tests for cattle is still very limited. The seed stock sector does appear to be one portion of the beef industry that is widely adopting genomic tools, though there is not a reliable estimate of the extent. Vestal et al. (2013) find it is not uncommon for bull sale catalog entries to provide genomic test results alongside other standard information about the expected genetic merit of sires—called Expected Progeny Difference (EPD). (See Box) Interestingly, these researchers found that adding DNA test information did not result in higher bull prices in their study, but they anticipate prices will respond as buyers become better at interpreting genomic test results.
An estimate of expected difference in the performance of future progeny for a particular characteristic or trait based on data from ancestors of a particular animal, the sire or dam in question, and the records of their progeny. For example, records on weaning weights are used to calculate the expected progeny difference for weaning weights if a particular bull is used. This is often expressed as a plus or minus in pounds. A bull with a weaning weight EPD of +31, means that calves from this bull are expected to be 31 pounds heavier at weaning compared to calves from an average sire. Often, EPDs are reported with a projection of accuracy. The closer to one, the more accurate the EPD is in predicting impact of using a particular sire for a particular trait. The EPD is based on past records of animals and does not use genetic information from DNA.
Genomics uses DNA marker tests to identify specific regions of chromosomes where genes are located that affect a particular trait. Many traits have multiple genes that affect them. A DNA test is performed to see if a particular animal has favorable combinations of alleles affecting the trait of interest. This information must be combined with information on the percentage of genetic variation explained by the test, accuracy of the test, data related to differences in performance, and heritability of the trait to predict a difference for a trait from a particular parent. Thus, genomics alone is not a predictor of performance, but improvement in animal and herd performance can be enhanced when combined with EPDs as genetic marker information can allow for early prediction of genetic merit for an animal.
As buyers of DNA-tested bulls, cow-calf ranch operators may be already benefiting from genomic information; however, there’s little evidence about use of genomic tools on the ranch itself. At a recent meeting with some Wyoming stock growers, most said they were “not very knowledgeable” about today’s genomic tools, most had never used them, and most identified testing costs as a barrier to adoption. Nonetheless, most participants said they are interested in potentially using DNA tests to better predict several economically relevant traits, especially reproductive performance, calving ease, and feed conversion efficiency. They were also interested in how DNA testing might help them decide which heifers—cows that have not yet had calves—to retain for breeding. These responses make good sense because cow-calf operators make more money if they lose fewer calves at birth, can increase the number of calves weaned per cow in the herd, expand the productive life of a cow or bull, or reach a desired weaning weight with less feed. These are traits over which the rancher may have some control, and even more so with the use of accurate, affordable DNA tests (Mitchell et al., 2009).
Beyond the ranch gate, the benefits of genomic tools are the subject of a small number of studies. We can imagine DNA tests might enhance the potential for a feedlot operator to contract with cow-calf operators who can verify their animals will achieve slaughter weight more quickly in the feedlot environment, or exhibit more valuable carcass traits, or be less likely to be susceptible to feedlot illnesses such as Bovine Respiratory Disease. DeVuyst et al. (2011) assessed the benefits of Neogen’s Igenity panel test, which, as table 1 shows, provides scores for a variety of growth and carcass traits of interest to feedlot operators, including average daily gain (ADG), marbling, rib-eye area, tenderness, fat thickness, and USDA Yield grade. They found statistically significant but somewhat low correlations between the Igenity scores and the observed values of these traits in the study animals. In addition, they found that of the panels analyzed only the marbling score contributed a net positive financial return. Thompson et al. (2014) evaluated the value to feedlot operators of using comprehensive tests with information on seven economically relevant traits to manage and sort cattle already in the feedlot. The authors found that marbling and ADG panel scores resulted in higher feed lot profits, but the cost of testing was larger than the associated returns.
Further downstream, genomic tools may increase interest by restaurateurs and retail chains in contracting directly with commercial beef producers. Such arrangements do happen now, as with Nolan Ryan All Natural Beef, but they are not the industry norm. DNA tests may hasten the trend toward such arrangements by making it easier for sellers to guarantee a particular quality of end product or for buyers to find it. Some companies, no doubt, will see potential value in controlling genomics from breeding to the dinner plate.
In addition to needing to understand the incentives for adoption by each industry segment, we will need to also assess how benefits of adoption are likely to be shared along the supply chain. As an example, Weaber and Lusk (2010) attempted this when they examined how using genomic tools to select breeding stock for meat tenderness could play out in changes in market prices and quantities throughout chain. They considered both consumer demand for tenderness and genetic testing costs, and concluded substantial economic benefits would be shared among industry participants over 20 years.
Borrowing a page from the history of genetic engineering of plants, consumer perceptions of genomic technologies for beef cattle also need to be studied sooner rather than later. If we had to predict, we would guess that interested consumers will “like” the use of genomic tools to select for some traits—for example, calving ease—but might “dislike” it for others. Although genomic prediction tools are not intended for the creation of “frankencows,” some consumers will fear that’s next. Ellen Goddard of the University of Alberta and colleagues in Europe are currently undertaking surveys of consumer perceptions in Canada and Europe (Goddard, 2013), but so far as we know no such efforts are underway in the United States.
USDA puts priority on “animal breeding, genetics, and genomics” research because “dramatic improvements in yields of animal protein are crucial in meeting the ever-increasing food needs in the United States and around the world” (USDA-NIFA, 2016a). According to experts, advances in genomics have the potential to bring about significant gains in efficiency of beef production (Goddard, 2012). Genomic science continues to advance and SNP studies for beef cattle to multiply. Meanwhile only a handful of economic analyses have assessed the profitability of DNA testing of cattle. To understand implications for food security, we need a good grasp of the status of genomic predictions of productivity and cost-reducing traits, the ability of commercial tests to deliver this information accurately, incentives for adoption in the commercial cow-calf sector, and the price and quantity effects of incorporating selection for such traits into the genetic stock.
Economic analyses could help USDA prioritize investments in genomic research on economically relevant traits: where’s the biggest expected bang for the buck? In Australia, economists have used a full beef industry model to assess returns—and their distribution across industry segments—to past genetic improvements (Farquharson et al., 2003; Zhao et al., 2000), and various R&D impact tools to calculate benefit-cost ratios for new genetic technologies developed by the Cooperative Research Center for Beef Genetic Technologies (Griffith and Burrow, 2015). Working in tandem with genomics experts, who can tell us more about the potential timeline and likelihood of development of genomic advancements, economists could evaluate which genomic advancements offer the most promise in terms of both net economic benefits and successful development. Economic analyses could also inform regulatory questions that may emerge down the road, such as the benefits of public or third-party validation services for beef genomic tools (Van Eenennaam et al., 2007). If you buy that 30 and Daisy test kit, you’ll want to know what you’re getting and whether it’s worth the cost.
Abo-Ismail, M.K., G. Vander Voort, J. Squires, K. Swanson, I. Mandell, X. Liao, P. Stothard, S. Moore, G. Plastow, and S. Miller. 2014. “Single Nucleotide Polymorphisms for Feed Efficiency and Performance in Crossbred Beef Cattle.” BMC Genetics 15:14.
AgriCultured. 2016. Certified Angus Beef. Available online: http://agricultured.org/certified-angus-beef/).
Bongiorni, S., G. Mancini, G. Chillemi, L. Pariset, and A. Valentini. 2012. “Identification of a Short Region on Chromosome 6 Affecting Direct Calving Ease in Piedmontese Cattle Breed.” PLoS ONE 7(12): e50137.
Field, T.G. 2007. Beef Production and Management Decisions. New Jersey: Pearson Prentice Hall.
Garrick, D.J. 2011. “The Nature, Scope and Impact of Genomic Prediction in Beef Cattle in the United States.” Genetics Selection Evolution 43: 2011.
Goddard, M.E. 2012. “Uses of Genomics in Livestock Agriculture.” Animal Production Science 52(3): 73-77.
Matukumalli, L.K., C.T. Lawley, R.D. Schnabel, J.F. Taylor, M.F. Allan, M.P. Heaton, J. O’Connell, S.S. Moore, T.P.L. Smith, T.S. Sonstegard, and C.P. Van Tassell. 2009. “Development and Characterization of a High-density SNP Genotyping Assay for Cattle.” PLoS ONE 4:e5350.
Meuwissen, T., B. Hayes, and M. Goddard. 2016. “Genomic Selection: A Paradigm Shift in Animal Breeding.” Animal Frontiers 6(1): 6-14.
Olmstead, A.L. and P.W. Rhode. 2008. “Creating Abundance: Biological Innovation and American Agricultural Development.” New York: Cambridge University Press.
Ossola, A. 2015. “I Got My Genes Tested. Should You?” Popular Science. Available online: http://www.popsci.com/i-tested-my-genes
Pollak, E.J., G.L. Bennett, W.M. Snelling, R.M. Thallman, and L.A. Kuehn. 2012. “Genomics and the Global Beef Cattle Industry.” Animal Production Science 52: 92-99.
Seidel, G.E. 2009. “Brief Introduction to Whole-genome Selection in Cattle Using Single Nucleotide Polymorphisms.” Reproduction, Fertility, and Development 22(1): 138-144.
Spangler, M. 2011. “Genetics of Beef Cattle: Moving to the Genomics Era.” Department of Animal Science, University of Nebraska-Lincoln.
Spangler, M. 2011. “Integration of Genomic Information into Genetic Evaluation.” 2011 NCBA Cattlemen’s College, Department of Animal Science, University of Nebraska-Lincoln.
The Bovine HapMap Consortium. 2009. “Genome-Wide Survey of SNP Variation Uncovers the Genetic Structure of Cattle Breeds.” Science 324 (5926): 528-532.
U.S. Department of Agriculture. National Institute of Food and Agriculture (USDA-NIFA). 2016a. Animal Breeding Genetics and Genomics. Available online: http://nifa.usda.gov/program/animal-breeding-genetics-and-genomics
U.S. Department of Agriculture. National Institute of Food and Agriculture (USDA-NIFA). 2016b. Research, Education, and Economics Information System. Project Search. Available online: http://portal.nifa.usda.gov/enterprise-search/
Van Eenennaam, A. 2010. “DNA-Based Biotechnologies.” In National Beef Cattle Evaluation Consortium, Sire Selection Manual, 2nd edition, pp. 68-78. Available online: http://www.nbcec.org/producers/sire.html
Van Eenennaam, A.L., J. Li, R. M. Thallman, R. L. Quaas, M. E. Dikeman, C. A. Gill, D. E. Franke and M. G. Thomas. 2007. “Validation of commercial DNA tests for quantitative beef quality traits.” Journal of Animal Science 85:891-900.
Alford, A.R., W.A. McKiernan, L.M. Cafe, P.L. Greenwood, and G.R. Griffith. 2009. “The Economic Effects of Using Heterozygotes for a Non-functional Myostatin Mutation within a Commercial Beef Production System.” NSW Department of Primary Industries Economic Research Report No. 42, Armidale, Australia.
DeVuyst, E.A., J.R. Bullinger, M.L. Bauer, P.T. Berg, and D.M. Larson. 2007. “An Economic Analysis of Genetic Information: Leptin Genotyping in Fed Cattle.“ Journal of Agricultural and Resource Economics 32(2): 291-305.
DeVuyst, E.A., J.T. Biermacher, J.L. Lusk, R.G. Mateescu, J.B. Blanton Jr., J.S. Swigert, B.J. Cook, and R.R. Reuter. 2011. “Relationships Between Fed Cattle Traits and Igenity Panel Scores.” Journal of Animal Science, Vol. 89: 1260-1269.
Farquharson, R.J., G.R. Griffith, S.A. Barwick, R.G. Banks, and W.E. Holmes. 2003. “Estimating the Returns from Past Investment into Beef Cattle Genetic Technologies in Australia.” Economics Research Report No. 15, NSW Agriculture, Armidale, Australia.
Goddard, E. 2013. “Canadian Views on Livestock Genomics and Public Investment in Genomic Technologies.” Presented at AARES 57th Annual Meeting, Sydney, NSW.
Griffith, G.R. and H.M. Burrow. 2015. “The Value of Research: Using the IMPACT TOOL to Evaluate Realized and Anticipated Benefits of the Cooperative Research Centre for Beef Genetic Technologies.” Animal Production Science 55: 133-144.
Mintert, J., T.C. Schroeder, G.W. Brester, D. Feuz. 1997. “Beef Industry Challenges and Opportunities.” In Managing for Today’s Cattle Market and Beyond. Western Extension Marketing Committee. Available online: http://marketing.uwagec.org/MngTCMkt/
Mitchell, J., E.A. DeVuyst, M.L. Bauer, and D.L. Larson. 2009. “Cow-calf Profitability and Leptin Genotyping.” Agricultural Economics 40:113-118.
Norwood, B. and J.L. Lusk. 2008. Agricultural Marketing and Price Analysis. New Jersey: Pearson Prentice Hall.
Office of Science and Technology Policy (OSTP). 2016. STAR METRICS. Available online: https://www.starmetrics.nih.gov/
Schaffer, J. 2015. Thesis proposal. Department of Agricultural and Applied Economics, University of Wyoming, Laramie, WY.
Thompson, N.M., E.A. DeVuyst, B.W. Brorsen, and J.L. Lusk. 2014. “Value of Genetic Information for Management and Selection of Feedlot Cattle.” Journal of Agricultural and Resource Economics 39(1): 139-155.
Van Eenennaam, A.L. and D.J. Drake. 2012. “Where in the Beef-Cattle Supply Chain Might DNA Tests Generate Value?” Animal Production Science 52: 185-196. Available online: http://www.gbcbiotech.com/bovinos/aplicaciones/trazabilidad/Where%20in%20the%20beef-cattle%20supply%20chain%20might%20DNA%20tests%20generate%20value.pdf
Vestal, M.K., J.L. Lusk, E.A. DeVuyst, and JR. Kropp. 2013. “The Value of Genetic Information to Livestock Buyers: a Combined Revealed, Stated Preference Approach.” Agricultural Economics 44: 337-347.
Weaber, R.L. and J.L. Lusk. 2010. “The Economic Value of Improvements in Beef Tenderness by Genetic Marker Selection.” American Journal of Agricultural Economics 92(5): 1456-1471.
Zhao, X., J.D. Mullen, G.R. Griffith, W.E. Griffiths, and R.R Piggott. 2000. “An Equilibrium Displacement Model of the Australian Beef Industry.” NSW Agriculture Economic Research Report No. 4, Orange, Australia.